The main steps of the algorithm are:
2a. Each PDB file is considered internally delimited by both "END" and "ENDM" records. This allows processing of multiple models in NMR files, and of concatenated PDB files. A command line option controls whether just the first or all of the models/structures within a file are processed.
2b. All atom records containing a character other than " ", "A" or
"1" in the alternate location column (column 17) were ignored.
The convention on alternate locations is that the column be blank if the
atom is unambiguously located, or contain the sequence "A", "B", "C" etc..
for each potential conformation for ambiguous atoms. Some PDB files use
digits instead of letters, for example pdb code 1icn.
2c. All atom records containing " Q" as the first two characters of the atom name were ignored. The element " Q" is commonly used in NMR processing to represent pseudo atoms used in refinement.
2d. The residue name "DUM" is officially used to denote dummy
atoms, such as in pdbcodes 1b3o and 1som, to
represent unexplained electron density. All atom records with residue
name "DUM" are ignored.
2e. All atoms with X, Y and Z co-ordinates equal 9999.000 are also ignored. These dummy atoms are created by XPLOR when the actual atom is not used in the structure refinement.
Occasionally even the above rules are insufficient. For example,
pdbcode 1a34 contains the same atom, O3* of U1D, twice.
First as atom serial number 2960 and again as 2973, both with identical
co-ordinates.
1e8b)
or incorrect atomic symbols (Nd in 1d00,
U in 1bt6, 1fgg, 1hh6,
3bcc).
Combined with the slow uptake of this field and the requirement to
process pre-version 2.0 files, its is better to just process the atom
name, which together with the residue name typically provide enough
contextual information to determine the atomic number.
3a. The atom name " UNK" is interpreted as an atom of atomic number, and is assigned element number 0, represented "*" in SMILES.
3b. If the first character of the atom name is blank, and the third character is lower case, the PDB file was generated by CONCORD and the atomic symbol is taken from the 2nd and 3rd characters. As PDB files are normally only uppercase, CONCORD's incorrect column alignment is easy to recognize.
3c. If the first character is blank, and residue name denotes
a hetero group that must have a prefix, and the third character is
"H", "C", "N", "O", "P" or "S", the appropriate atomic number is used.
If the third character is not one of these, the 2nd character is used
as the atomic symbol. The current hetero groups that require a prefix
are "GPC", "NAD" and "NDP" (which fixes 1rds).
3d. If the first character is blank, the second character is assumed to be the atomic symbol. If this character isn't recognized as an atomic symbol, and the third character contains "H", "C", "N", "O", "P" or "S", then the appropriate atomic number is used.
3e. If the first character is a digit, the second character is assumed to be the atomic symbol.
3f. If the first character is "H", and the residue is a
recognised amino acid, nucleic acid or special hetero group, then
the atom is assumed to be hydrogen. For remaining groups, if the
first two characters are a recognized atomic symbol, this is the
element, or hydrogen if the first two characters are not a recognized
atomic symbol. This fixes the Holmiums "Ho" and Mercuries "Hg" in
numerous files (including 1f85).
The recognized amino acid residues are "ACE", "ALA", "ARG", "ASN", "ASP", "ASX", "CYS", "FOR", "GLN", "GLU", "GLX", "GLY", "HIS", "HYP", "ILE", "LEU", "LYS", "MET", "PHE", "PRO", "PCA", "SER", "THR", "TRP", "TYR", "UNK" and "VAL".
The recognized nucleic acid residues are " A", " C", " G", " T", " U", " +U", " YG", "1MA", "1MG", "2MG", "5MC", "5MU", "7MG", "H2U", "M2G", "OMC", "OMG" and "PSU".
The recognized special hetero groups are "101", "12A", "1AR", "1GL",
"2AS", "2GL", "3AA", "3AT", "3DR", "3PO", "6HA", "6HC", "6HG", "6HT",
"A26", "AA6", "ABD", "AC1", "ACO", "AIR", "AMU", "AMX", "AP5", "AMG",
"APU", "B9A", "BCA", "BNA", "CAA", "CBS", "CGS", "CMC", "CND", "CO8",
"COA", "COF", "COS", "DCA", "DGD", "FAB", "FAD", "FAG", "FAM", "FDA",
"GPC", "IB2", "NAD", "NAH", "NAI", "NAL", "NAP", "NBD", "NDP", "PAD",
"SAD", "SAE", "T5A", "tRE", "UP5" and "ZID". Although it shouldn't
be required including "BU1" fixes 1bk9.
3g. If the first character is one of the symbols """, "'" or *" then the second character is treated as the atomic symbol.
3h. If the residue name indicates a hetero group that uses
an suffix, the the first character of the atom name is treated as
the atomic symbol. The current hetero groups that require a suffix
are "AGF", "COT" amd "FVF" (which fixes 1cjw).
3i. If the residue name is one of the special hetero groups listed in 3f, the second character of the atom name is treated as the atomic symbol.
3j. The default, all remaining cases, treats the first two characters of the atom name as the atomic symbol. If the first two characters aren't a valid element, then the second character is treated as the atomic symbol.
3k. An exception to the above rule 3j is that "Nd"
(atomic number 60) doesn't occur in any of the amino acids listed
in section 3f, and occurances are corrected to nitrogen.
This fixes pdbcode 1d00.
3l. Finally, exceptions to all of the above rules are
handled explicitly. Currently, the only exception is the atom
name "NSE1" which occurs in residues "SAD" and "SAE". This atom
name actually represents a selenium atom, encoded in the 2nd and
3rd columns! (which handles 1adg and 1b3o).
4a. Metal atoms are prevented from forming covalent bonds. It appears that software used for protein x-ray crystallography (such as XPLOR and CNS) are poorly parameterized for inorganic compounds when compared with small molecule crystallography software (such as SHEL-X). The false positive rate when using proximity based connectivity perception is much higher in PDB than in the Cambridge Structural Database (CSD).
Unfortunately, this constraint means that algorithm fails for some of
the organo-metalics in PDB, including the ferrocene in 1a3l
and the uridine vanadate in 6rsa.
There is a remarkable diversity of metalic elements in the PDB:
| Element | Symbol | Number | Example PDB Code |
| Lithium | Li | 3 | 1e5k
|
| Beryllium | Be | 4 | 4ukd
|
| Magnesium | Mg | 12 | 1dpl
|
| Aluminium | Al | 13 | 1xlm
|
| Argon | Ar | 18 | 1c6i
|
| Vanadium | V | 23 | 1dkt
|
| Chromium | Cr | 24 | 1cf6
|
| Manganese | Mn | 25 | 1nls
|
| Cobalt | Co | 27 | 1b6a
|
| Copper | Cu | 29 | 1mfm
|
| Zinc | Zn | 30 | 1dzv
|
| Gallium | Ga | 31 | 1cfw
|
| Arsenic | As | 33 | 3cao
|
| Krypton | Kr | 36 | 1c6g
|
| Rubidium | Rb | 37 | 460d
|
| Strontium | Sr | 38 | 434d
|
| Yttrium | Y | 39 | 1dde
|
| Molybdenum | Mo | 42 | 1g8k
|
| Silver | Ag | 47 | 1aoo
|
| Cadmium | Cd | 48 | 1mfm
|
| Indium | In | 49 | 1ind
|
| Antimony | Sb | 51 | 1f48
|
| Tellurium | Te | 52 | 1el7
|
| Xenon | Xe | 54 | 1c6e
|
| Caesium | Cs | 55 | 1av2
|
| Barium | Ba | 56 | 284d
|
| Lanthanum | La | 57 | 1djg
|
| Cerium | Ce | 58 | 1ak8
|
| Samarium | Sm | 62 | 1a3c
|
| Europium | Eu | 63 | 1qsl
|
| Gadolinium | Gd | 64 | 2hhm
|
| Terbium | Tb | 65 | 1ncz
|
| Holmium | Ho | 67 | 1psr
|
| Ytterbium | Yb | 70 | 2bop
|
| Lutetium | Lu | 71 | 1e8x
|
| Tungsten | W | 74 | 6fit
|
| Rhenium | Re | 75 | 1b0q
|
| Osmium | Os | 76 | 1qa6
|
| Iridium | Ir | 77 | 1c1k
|
| Platinum | Pt | 78 | 1qbi
|
| Gold | Au | 79 | 1a8d
|
| Mercury | Hg | 80 | 1cc8
|
| Thallium | Tl | 81 | 1fpj
|
| Lead | Pb | 82 | 1xxa
|
| Uranium | U | 92 | 1b5j
|
4b. Two atoms are considered bonded if they are closer than the sum of their covalent radii plus a small tolerance. The values of covalent radii are those published by the Cambridge Crystallographic Data Center and are given in the table below. Although the CCDC recommend a tolerance of 0.4 Angstroms, the current algorithm uses the larger value 0.45 Angstroms used by Baber and Hodgkin, and by Hendlich et al.. This value is lower than the 0.56 Angstrom tolerance used by the molecular graphics program RasMol. A lower bound of 0.4A for bond lengths is also imposed.
| Element | Symbol | Number | Radius (A) |
| Hydrogen | H | 1 | 0.23 |
| Boron | B | 5 | 0.83 |
| Carbon | C | 6 | 0.68 |
| Nitrogen | N | 7 | 0.68 |
| Oxygen | O | 8 | 0.68 |
| Fluorine | F | 9 | 0.64 |
| Silicon | Si | 14 | 1.20 |
| Phosphorus | P | 15 | 1.05 |
| Sulfur | S | 16 | 1.02 |
| Chlorine | Cl | 17 | 0.99 |
| Arsenic | As | 33 | 1.21 |
| Selenium | Se | 34 | 1.22 |
| Bromine | Br | 35 | 1.21 |
| Telurium | Te | 52 | 1.47 |
| Iodine | I | 53 | 1.40 |
4c. Bonds can only be formed between atoms with the same chain
identifier and that are not separated by a TER (chain terminator)
record in the PDB file. Unfortunately the TER record in PDB file
1atl is incorrectly placed, cleaving a methyl group.
4d. All solvent atoms are prevented from forming covalent bonds other than to the same residue. Close contacts with waters account for a frequent source of false positive bonds. Recognizing the residue names for water ("HOH", "H20", "WAT", "TIP", "SOL", "DOD" and "D20") and for other solvents ("EOH", "MOH", "PER", "PO4", "SO4" and "SUL"), and explicitly excluding these atoms from inter-residue bonding improves the resuting topologies.
4e. Any hydrogens that are two or more connected by the above procedure are then fixed. All its bonds, except the one to the closest non-hydrogen atom in the same residue, are deleted.
4f. Occasionally, atoms with identical co-ordinates are present in a PDB file. If two atoms in different PDB groups have the same co-ordinates and are bonded to a common third atom, all bonds between the these two groups are broken. If two atoms with the same co-ordinate remain bonded to a common third atom, the second atom is deleted.
4g. If any atom is bonded to more than a maximum number of
neighbors for that element, then bonds between the PDB group containing
that atom and groups connected via that atom are broken. This handles
multiple occupancy problems such as 1abe. The maximum
neighbor count for each element is four unless tabulated below.
| Element | Symbol | Number | Max Neighbors |
| Hydrogen | H | 1 | 1 |
| Boron | B | 5 | 3 |
| Oxygen | O | 8 | 2 |
| Fluorine | F | 9 | 1 |
| Bromine | Br | 35 | 1 |
| Iodine | I | 53 | 3 |
1ela for
example) and not always honored (pdbcodes 1dxd,
1qlb, 5ana and 1sdg).
5a. All connected components containing more than 100 heavy atoms are considered to be protein (or nucleic acid). If the entire component originated from ATOM records, it is ignored. Otherwise, all ATOM atoms are deleted, except for those covalently bonded directly to HETATM atoms. The atoms are converted to an asterisk to indicate an attachment point, and bonds between pairs of asterisks are deleted. This should have the effect of eliminating all large proteins and nucleic acids, after cleaving all covalently bound ligands and post-translational modifications.
5b. The next step is a fragment size filter. All remaining connected components (after step 5a) are ignored if they contain more than 100 or less than six heavy atoms (typically metals, waters, sulphates and phosphates).
5c. The final optional step is to remove all protein and nucleic
acid fragments. These are all connected components (after 5b)
that contain no HETATM records. If the PDB file contained a connected
component of more than 100 atoms (a protein or nucleic acid), an exception
is made for components whose chain identifiers occured in no other
components. This correctly identifies peptide ligands (such as chain
"I" in 4hvp, or chain "C" in 1hsl).
Unfortunately, the lysine in 1lst is still not correctly
perceived as a ligand.
6a. Initial assignment of hybridization state is on the basis of average bond angle. Two connected atoms with a bond angle of greater than 155 degrees are typed as sp-hybridized. Remaining atoms with an average bond angle greater than 115 degrees are typed as sp2-hybridized and those with an average bond angle less than or equal to 115 degrees as sp3-hybridized.
6b. Average bond angles are unable to descriminate the correct hybridization state of two connected atoms in five membered rings. For example, in an aromatic ring such as pyrole or furan, the sp2 carbons have bond angles near 108 degrees! A second pass sets the hybridization of all two connected atoms in a five membered ring, as sp2-hybridized when the average in-ring torsion is less than 7.5 degrees. A similar planarity test is used for six membered rings, where a 12 degree threshold is used.
6c. A final 'antialiasing' pass is used to detect and correct misassigned hybridization states. If an sp-hybridized atom doesn't have a sp-hybridized or terminal neighbor with unfilled valence, it is reassigned as sp2-hybridized. Similarly, if an sp2-hybridized atom doesn't have a sp2-hybridized or terminal neighbor with unfilled valence, it is reassigned as sp3-hybridized.

Because the recognition is applied after hybridization state assigment,
the a patterns can make use of geometry at each pattern position.
Typically, however, hybridization information is only used for ring
atoms. This correctly handles distorted carboxylic acid groups (such
as in 1tdr) but also correctly handles difficult systems
like 2ada.
When multiple patterns are applicable heuristics based upon atomic electronegativity are used to place the double bond. A double bond to a terminal oxygen is chosen over a double bond to a terminal sulphur. For guanadine groups, if the central carbon is in a ring a ring nitrogen is chosen over a non-ring nitrogen, otherwise a terminal nitrogen is chosen over a non-terminal nitrogen.

Under each of the atom types above are listed the number of electrons each contributes to the ring for the Huckel 4n+2 aromaticity calculation. Carbons typically contribute one, except when doubly bonded to a terminal oxygen atom. Oxygen, Sulphur and Selenium contribute two. Pyridine and N-oxide nitrogens contribute one, and pyrole nitrogens contribute two.
At this point, there are two ambiguous cases. The first is that a carbon of unfilled valence can double bond to the oxygen or the ring, resulting in zero or one electrons respectively. The second is that a two connected nitrogen of unfilled valence could be double bonded to the ring to become pyridine-like, or gain an implicit hydrogen to become pyrole-like, contributing one and two electrons respectively.
The heuristic used by the algorithm is that a sp2-hybridized ring should be made aromatic if possible. This is done in the following steps.
8a. First, a test is applied to the ambiguous cases (*-[C](O)-* and *-[N]-*) to see whether they can be resolved by their neighbours. If both neighbours have full valences or incident multiple bonds, both ring bonds must be single and their unique types are reassigned (*-C(=O)-* and *-[NH]-* respectively).
8b. If the count of electrons in the ring modulo four is one, and the ring contains an ambiguous nitrogen, it is retyped pyrole-like (with an implicit hydrogen).
8c. If the count of electrons in the ring modulo four is three, and the ring contains an ambiguous carbon, it is retyped with an exo double bond to the oxygen.
8d. If the count of electrons in the ring modulo four is three, and the ring contains a uncharged pyrole-like nitrogen, the nitrogen is charged.
8e. If, after all the above reassignments, the count of electrons in the ring modulo four is two, the Huckel condition, all the atoms and bonds are marked as potentially aromatic.
After the above processing has been applied to all rings in a molecule, the molecule is passed to a kekule form assignment routine that attempt to provide a kekule form of alternating single and double bonds. The actual kekule form assignment algorithm is complex and described in detail elsewhere (parts of the algorithm were presented in the tautomerism talk at EuroMug99).
9a. The first stage of final bond order assignment is to mark all ring bonds between sp2-hybridized atoms as aromatic, and call the kekule assignment algorithm a second time. This should assign alternating single and double bonds to conjugated ring systems including aromatic rings with more than seven atoms.
9b. The next step is to process any unsatisfied sp2-hybridized atom by checking each neighboring atom for a terminal oxygen, and when found testing the bond length against the table of bond lengths below. This pass preferencially selects the keto over the enol forms of conjugated molecules.
9c. The very last step is to check each bond using distance criteria. Only bonds between atoms with unfilled valences and without an incident multiple bond are considered. Bonds between sp-hybrized atoms or between sp-hybridized and terminal atoms are tested against triple bond lengths, and bonds between sp2-hybridized atoms or between sp2-hybridized atoms and terminal atoms are tested against double bond lengths. Bonds between a pair of terminal atoms are tested against both double and triple bond lengths.
| Bond | Distance | Example |
| C#C | 1.25 | 1nco
|
| C#N | 1.22 | 2cgr
|
| C=C | 1.38 | 1rbp
|
| C=O | 1.28 | 3cpp
|
| C=S | 1.70 | 8cpp
|
| N=N | 1.32 | 1srj
|
9d. All remaining unfilled valences are filled with implicit hydrogen atoms. The atomic valences are the standard values used by the Daylight toolkit.
The most common ligands are heme and cytochrome (456+242 occurences), followed by N-acetyl-D-glucosamine "NAG" (341 occurences), followed by glycerol "GOL" (222 occurences). 2426 ligands occur only once, with the remaining 1075 occuring multiple times (135 >10, 39 >25, 23 >50 and 12 >100).
On test set 1 of protein ligands from the review by Ricketts et al.. The algorithm gets all 18 of the ligands correct. Back in 1996 the author did a review of existing techniques on a subset of 17 of these files. The results are shown in the table below. The column labelled "All Single" shows the structures correct when all bonds are left single. The "Covalent" method (inspired by Pauling's nature of the chemical bond) assigns a triple bond to lengths less than 81% of the sum of the attached covalent radii, a double bond between 81% and 87%, and a single bond above 87%. The "SMARTS" method used simple pattern matching of terminal groups (similar to the functional group recognition in section 7). The "Cambridge" and "Oxbridge" algorithms are based on the algorithm of Baber and Hodgkin. The "Cambridge" column are the results using just bond lengths for CSD, and "Oxbridge" is the full algorithm using bond lengths and bond angles. The "IDATM" column represents the algorithm of Meng and Lewis as encoded in Babel v1.4. The "COBRA" column is Andrew Leach's perception code as used by Oxford Molecular's COBRA package. The "BONDAGE" column contains the results of algorithm by Blaney, Dixon and Swanson in the DGEOM95 package.
| No. | PDB Code | All Single | Covalent | SMARTS | Cambridge | Oxbridge | IDATM | COBRA | Bondage | This work |
| 1 | 1cla | N | N | N | N | N | N | N | N | Y |
| 2 | 2aat | N | N | N | N | N | N | N | N | Y |
| 3 | 2gbp | Y | Y | Y | Y | Y | Y | Y | Y | Y |
| 4 | 3cpp | N | Y | N | Y | Y | Y | N | Y | Y |
| 5 | 2trm | N | N | N | N | Y | N | N | N | Y |
| 6 | 3ptb | N | N | N | N | Y | N | N | Y | Y |
| 7 | 5xia | Y | Y | Y | Y | Y | Y | Y | Y | Y |
| 8 | 4xia | Y | Y | Y | Y | Y | Y | Y | Y | Y |
| 9 | 8ldh | N | N | Y | Y | N | N | Y | Y | Y |
| 10 | 8atc | N | N | N | N | N | N | Y | N | Y |
| 11 | 8rsa | N | N | N | N | Y | Y | N | N | Y |
| 12 | 1fcb | N | N | N | N | N | N | N | N | Y |
| 13 | 1fx1 | N | N | N | N | N | N | N | N | Y |
| 14 | 1gox | N | N | N | N | N | N | N | N | Y |
| 15 | 2dhf | N | N | N | N | N | N | Y | N | Y |
| 16 | 4dfr | N | N | N | N | N | N | N | N | Y |
| 17 | 7dfr | N | N | N | N | N | N | N | N | Y |
| Totals | 3 | 4 | 4 | 5 | 7 | 5 | 6 | 6 | 17 |
Note these results are slightly biased as results were used in the design and development of the algorithm presented above. Also note that the table is actually presented in approximate order of the quality of results, and although Baber and Hodgkin's Oxbridge algorithm looks impressive, for the structures is got, it performed far worse than the methods to its right.
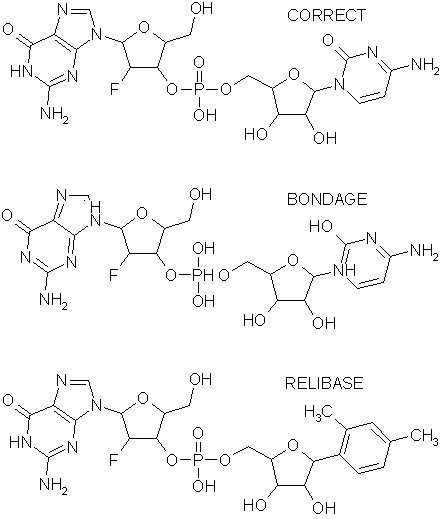
On the DockIt test set of 10 PDB files, the program gets all 10 ligands
correct. The original FORTRAN bondage algorithm gets 8/10, and CCDC's
ReliBase, which is hand curated, also gets 8/10. ReliBase doesn't consider
the peptide inhibitor of HIV protease in 4hvp to be a ligand,
and gets the element typing in 1rds wrong.
Hendlich reports in his paper, that the BALI program gets the biotin
in 1bib wrong due to an abnormal bond length of 1.133A
which is typed as a triple bond. The described algorithm get 1bib
correct as a triple bond requires supporting sp-hybridization evidence.
Finally the 133 pdb codes below are taken from the extended GOLD benchmark suite of structures. The "This" column represents the algorithm described here, the "Gold" column represents the "ligand.mol2" structures distributed with gold, and the "Reli" column represents the structures in Relibase v4.0.
| No. | PDB Code | This | Gold | Reli |
| 1 | 1aaq | Y | Y | Y |
| 2 | 1abe | Y | Y | Y |
| 3 | 1acj | Y | N | Y |
| 4 | 1ack | Y | Y | - |
| 5 | 1acm | Y | Y | ? |
| 6 | 1aco | Y | Y | ? |
| 7 | 1aec | Y | N | ? |
| 8 | 1aha | Y | Y | Y |
| 9 | 1apt | Y | Y | N |
| 10 | 1ase | Y | N | N |
| 11 | 1atl | N | Y | N |
| 12 | 1azm | Y | Y | Y |
| 13 | 1baf | Y | Y | Y |
| 14 | 1bbp | N | Y | Y |
| 15 | 1blh | Y | Y | Y |
| 16 | 1bma | Y | Y | Y |
| 17 | 1byb | Y | Y | Y |
| 18 | 1cbs | Y | Y | Y |
| 19 | 1cbx | Y | Y | Y |
| 20 | 1cdg | Y | Y | N |
| 21 | 1cil | Y | Y | Y |
| 22 | 1com | Y | N | Y |
| 23 | 1coy | Y | Y | Y |
| 24 | 1cps | Y | Y | Y |
| 25 | 1ctr | Y | Y | Y |
| 26 | 1dbb | Y | Y | Y |
| 27 | 1dbj | Y | Y | Y |
| 28 | 1did | Y | Y | Y |
| 29 | 1die | Y | Y | Y |
| 30 | 1dr1 | Y | Y | Y |
| 31 | 1dwd | Y | Y | Y |
| 32 | 1eap | Y | Y | Y |
| 33 | 1eed | Y | ? | Y |
| 34 | 1epb | Y | Y | Y |
| 35 | 1eta | Y | Y | Y |
| 36 | 1etr | Y | N | Y |
| 37 | 1fen | N | Y | Y |
| 38 | 1fkg | Y | Y | Y |
| 39 | 1fki | Y | Y | Y |
| 40 | 1frp | Y | Y | Y |
| 41 | 1ghb | Y | Y | Y |
| 42 | 1glp | Y | N | ? |
| 43 | 1glq | Y | Y | N |
| 44 | 1hdc | N | Y | Y |
| 45 | 1hdy | Y | Y | Y |
| 46 | 1hef | Y | Y | Y |
| 47 | 1hfc | Y | Y | Y |
| 48 | 1hri | Y | Y | Y |
| 49 | 1hsl | Y | Y | Y |
| 50 | 1hyt | Y | N | Y |
| 51 | 1icn | Y | N | ? |
| 52 | 1ida | Y | Y | Y |
| 53 | 1igj | Y | ? | Y |
| 54 | 1imb | Y | Y | Y |
| 55 | 1ive | Y | Y | Y |
| 56 | 1lah | Y | Y | ? |
| 57 | 1lcp | Y | Y | ? |
| 58 | 1ldm | Y | Y | ? |
| 59 | 1lic | Y | N | Y |
| 60 | 1lmo | Y | Y | Y |
| 61 | 1lna | Y | Y | Y |
| 62 | 1lpm | Y | N | Y |
| 63 | 1lst | - | Y | - |
| 64 | 1mcr | Y | Y | Y |
| 65 | 1mdr | Y | Y | Y |
| 66 | 1mmq | Y | Y | Y |
| 67 | 1mrg | Y | Y | Y |
| 68 | 1mrk | Y | Y | Y |
| 69 | 1mup | N | Y | Y |
| 70 | 1nco | N | Y | Y |
| 71 | 1nis | Y | Y | ? |
| 72 | 1pbd | Y | Y | Y |
| 73 | 1pha | Y | Y | Y |
| 74 | 1phd | Y | Y | Y |
| 75 | 1phg | Y | Y | Y |
| 76 | 1poc | Y | Y | Y |
| 77 | 1rds | Y | Y | N |
| 78 | 1rne | Y | Y | Y |
| 79 | 1rob | Y | Y | Y |
| 80 | 1slt | Y | Y | Y |
| 81 | 1snc | Y | N | Y |
| 82 | 1srj | Y | Y | Y |
| 83 | 1stp | Y | Y | Y |
| 84 | 1tdb | Y | Y | Y |
| 85 | 1tka | N | Y | Y |
| 86 | 1tmn | N | Y | Y |
| 87 | 1tng | Y | Y | Y |
| 88 | 1tni | Y | Y | Y |
| 89 | 1tnl | Y | Y | Y |
| 90 | 1tph | Y | N | Y |
| 91 | 1tpp | Y | ? | Y |
| 92 | 1trk | N | N | Y |
| 93 | 1tyl | Y | Y | Y |
| 94 | 1ukz | Y | Y | Y |
| 95 | 1ulb | Y | Y | Y |
| 96 | 1wap | Y | Y | Y |
| 97 | 1xid | Y | Y | Y |
| 98 | 1xie | Y | Y | Y |
| 99 | 2ada | Y | Y | Y |
| 100 | 2ak3 | Y | Y | Y |
| 101 | 2cgr | Y | N | Y |
| 102 | 2cht | Y | Y | Y |
| 103 | 2cmd | Y | Y | ? |
| 104 | 2ctc | Y | Y | Y |
| 105 | 2dbl | N | Y | Y |
| 106 | 2gbp | Y | Y | Y |
| 107 | 2lgs | Y | Y | ? |
| 108 | 2mcp | Y | Y | ? |
| 109 | 2mth | Y | Y | - |
| 110 | 2phh | Y | Y | Y |
| 111 | 2pk4 | Y | Y | ? |
| 112 | 2plv | N | Y | Y |
| 113 | 2r07 | Y | Y | N |
| 114 | 2sim | Y | Y | Y |
| 115 | 2yhx | Y | Y | Y |
| 116 | 3aah | N | Y | - |
| 117 | 3cla | Y | Y | Y |
| 118 | 3cpa | Y | Y | Y |
| 119 | 3gch | N | ? | Y |
| 120 | 3hvt | Y | Y | Y |
| 121 | 3ptb | Y | Y | Y |
| 122 | 3tpi | Y | Y | ? |
| 123 | 4cts | Y | Y | ? |
| 124 | 4dfr | Y | Y | Y |
| 125 | 4est | N | ? | Y |
| 126 | 4fab | Y | N | Y |
| 127 | 4phv | Y | Y | Y |
| 128 | 5p2p | Y | Y | ? |
| 129 | 6abp | Y | Y | Y |
| 130 | 6rnt | N | Y | Y |
| 131 | 6rsa | N | ? | Y |
| 132 | 7tim | Y | Y | ? |
| 133 | 8gch | Y | Y | Y |
| Totals | 116 | 112 | 105 |
The files 1ack, 2mth and 3aah are
obsolete and have been replaced in the PDB by corrected versions.
Relibase consistently represents nitro groups incorrectly. The lysine
in 1lst is not perceived as a ligand in this work or
Relibase. The vanadium atom in 6rsa causes problems for
the described algorithm and GOLD (where it is replaced by a phosphorus).
1bbp is a particularly tricky chromophore. There's a
misplaced "TER" record in 1atl.