7. THOR - Chemical Database System
THOR (THesaurus Oriented Retrieval) is a chemical database system. That is, it is specifically designed to store and retrieve chemical information efficiently. Because THOR has built-in knowledge about chemical graph theory, it achieves high performance when storing chemical information. And more importantly, because it is specifically designed to store chemical information, it stores and retrieves it in ways that make sense to the chemist.
THOR's "language" is the common language of chemists. Users can store information using whatever nomenclature is appropriate or convenient; THOR understands the relationships between various nomenclatures and can retrieve a compound's data using any name ("identifier") that is known for the compound. For example, one might at different times choose to use SMILES, Wiswesser Line Notation (WLN), CAS numbers, local ID's, common, trade, and formal names. THOR allows chemical information to be referenced by an unlimited number of imprecise synonyms (e.g. states, mixtures, isomers, congeners, conformations), and maintains the relationships between each name, what is known about it, and how it is related to other names.
THOR makes no assumptions about the type or nature of the data that is to be stored, other than that it is information about chemical entities. Users can freely define their own datatypes -- descriptions of the data that are to be stored -- so any type or quantity of data can be stored. The definition of a datatype does not in itself cause any space to be used in a THOR database; space is allocated only when actual data are stored. Thus, hundreds or thousands of different types of data can reside in a single database efficiently.
7.1 Hash Table
Computer disks are designed to hold large amounts of information at relatively low cost, but they are slow at random-access operations - a typical disk can only make a few hundred random accesses per second. Thus, a database system that uses disk-resident databases should minimize the number of disk accesses it makes.
By using the unique SMILES of a molecule as the molecule's primary identifier (the TDT's "main topic"), THOR is able to eliminate all searching during data retrieval. All data are looked up directly in a hash table. This section describes, in an introductory fashion, how THOR's hash tables work.
Hashing begins with a hash function, h(K,N), which takes a string of characters, K, and converts it (via a pseudo-randomizing algorithm) into a number between zero and N-1. For example, given an identifier such as "Oc1ccccc1" that we want to find in the database, h("Oc1ccccc1",101) might produce the number 76.
The hash function has three noteworthy properties:- Given a large number of keys K1-Kn, the integers I1-In produced by h(Ki,N) will be evenly distributed in the range 0 to N-1.
- The hash function can be computed very quickly.
- Collisions are possible, where two different keys Ki and Kj both produce the same result I.
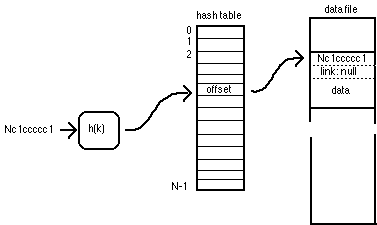
Hashing to access a non-colliding record
In any hash-table system, collisions are possible: two keys Ki and Kj can hash to the same value (that is, h(Ki,N) equals h(Kj,N). When this happens, a collision chain is formed in the database (shown in the illustration below). The hash table always contains the location of the head record, and that record's link holds the location of the previous record with the same h(K,N), and so on to the last record in the chain, which has a null location to indicate that the end of the chain has been reached.
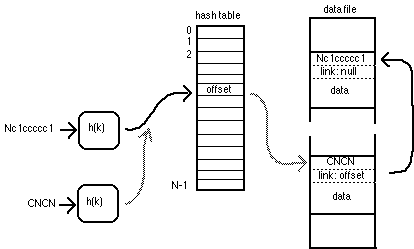
A collision-resolution chain for colliding records
Unless a database has zero or one records in it, there is a non-zero probability of collisions. As an analogy, imagine asking a random-number generator for 100 random numbers between 0 and 99. If the generator is truly random, one might expect roughly 70-80 unique numbers, and 20-30 duplicates. In a hash table where N=100, we can predict the same behavior when we store 100 records: there will be something like 20-30 collisions.
These collisions inevitably slow the data-access process down, since several disk accesses may be needed to find a record. In fact, at first glance one might guess that very long collision chains could form by random chance. Fortunately the odds are in our favor: The chances of very long chains forming are very remote, due to the statistics of random numbers.
More importantly, we can control these probabilities by setting the size of our hash table. If we know there are K records to be stored, a hash table with N=K/4 entries will give poor performance, one with N=K/2 entries will give reasonable performance, and one with N=K entries will give good performance.
THOR also has the ability to resize a hash table, because the data file and the hash table are completely separate, a hash table that is overloaded (K is much larger than N) can be discarded and a new, larger one built.
The most important feature of the hash-table mechanism is that data are accessed directly rather than by a time-consuming search. Hash tables exhibit what computer scientists call O(1) (pronounced "order one") performance: The access time of data doesn't depend on the size of the database. Looking at the examples above, we can see that the time required to look up any particular record is independent of how much data we have stored in the database. No matter how big the database becomes, the data access time only depends on three factors: how long it takes to compute h(K,N), how fast the disk is, and how many collisions we have on average. For a typical THOR system, h(k,N) takes a few microseconds, the average disk access is a few milliseconds, and if the hash table is properly sized, the average number of collisions is 1.2 to 1.5. Thus, access is roughly equal to the disk's access time.
The ability to use hash tables in THOR is directly related to the properties of SMILES. Unlike other identifiers used to name molecules, the unique SMILES is a fundamental property of the molecule, and there is only one unique SMILES for any particular molecule. This means that no matter how large a database grows, and no matter how many people enter data, all data for a particular molecule will be stored in the same record, and we can find that record by direct look-up rather than by searching.
7.2 Servers and Clients
The THOR system consists of two parts: servers and clients. The THOR server provides the ability to create, add data to, and read data from disk-based databases. The server allows many clients to read and write to the same database simultaneously, thus sharing the cost of the resource (disk) among many users. It can even provide database sharing across a wide geographical area.
In contrast, most of the complex aspects of THOR's operation, such as "standardizing" input file format, generating unique SMILES, merging input files, and so on, are handled at the client end. Many THOR client programs have a user interface of some sort. For example, the XVTHOR program uses X-Windows to provide a graphical display of data from a THOR database, and allows users to enter and edit chemical information; similarly, the 'sTHORman' program uses a "tty-style" user interface to manage THOR database. By contrast, the 'THORload' program has no user interface - it is "launched" and proceeds to load a database with no further user interaction.
7.3 Identifiers
THOR carefully distinguishes between identifiers and data. An
identifier is something about which you have data, or about which you could
have data. It is something to which you wish to attach information. As the
name implies, it identifies.
THOR is unique in its ability to use a wide variety of different identifiers to
access information, including imprecise synonyms, ambiguous names, isomeric and
tautomeric names, 3D conformations, trade names, common names, formal names,
and so forth. Some examples of common identifiers are SMILES, CAS number,
company IDs, IUPAC name, and Wiswesser Line Notation (WLN). The choice of
identifiers in THOR is up to the THOR manager; no restrictions are placed on
what constitutes a proper identifier, but the "philosophy" of how to choose
identifiers is important.
Frequently, identifiers are arbitrary names created by some convention or passed down through history. Examples of these arbitrary names are registrations numbers such as CAS, catalog numbers from vendors, and trivial names such as "morphine". Such identifiers carry no information about the chemical itself - the CAS number of 31604-28-1 is meaningless without a database. Likewise, "morphine" only has meaning because everyone knows what it is; the word itself has no chemical information.
But many identifiers do carry information about the chemical. Examples of this are IUPAC names, Wiswesser Line Notation (WLN), and SMILES; these all contain structural data. In other words, the name is both an identifier and contains chemical information. Since THOR is a database containing chemical information, we have to have a clear understanding about this dual role: that one thing can serve as both an identifier and as data.
It is rare for an identifier to name a single molecule with no possible variations. For example, 'dichlorobenzene' names three molecules: the meta-, ortho-, and para-substituted configurations; 'dichloroethylene' names the 1,1 as well as 1,2-cis and -trans configurations; the SMILES 'NC(CC)Cl' is a chiral structure with two possible configurations.
In contrast to the many other options, unique SMILES makes an ideal identifier for TDTs because of its special properties:
-
Unlike many identifiers, SMILES represents a fundamental property of a
molecule: Its bonds and atoms. Unlike common names, trade names, catalog
numbers and company stock numbers, SMILES is not arbitrary. Rather, it
represents information derived from the molecule itself. There is no
arbitrariness about a SMILES.
-
There no uncertainty about what is being represented; the
meaning of each SMILES is unambiguous. For example, although ClC=CCl
represents two molecules (cis- and trans-dichloroethene), it is clear that
this is the case. Given any molecule, one can write down its SMILES; given
any SMILES, one can tell if it represents a particular molecule. By contrast,
although virtually every chemist would interpret "1,2-dichloroethene" to
mean a mixture of the cis and trans configurations, there is nothing
requiring this interpretation except convention.
-
SMILES is understandable; most chemists learn basic SMILES in just a few
minutes.
-
SMILES can be read rapidly by a computer (this is not true of some line
notations; for example it is very difficult to write a program to read
Wiswesser Line Notation).
- Each molecule can be described by several SMILES, but the CANGEN algorithm will generate one special SMILES, the unique SMILES, for each molecule. The unique SMILES is universal: It is the same for every THOR database in the world. All THOR databases are intercompatible.
- The unique SMILES can be used to directly access THOR's records; no searching is necessary. This is discussed in more detail in the section on hash tables.
7.4 The THOR Data Tree
THOR uses a format with thesaurus orientation referred to as a THOR Data Tree or TDT in order to distinguish between identifiers and data associated with each identifier. To better understand this orientation, take a look at the traditional use of a thesaurus to capture relationships between a topic with one or more subtopics describing related ideas or concepts. Consider these two examples:
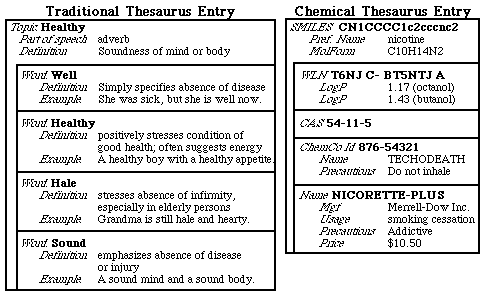
The above examples illustrate several important points about thesaurus entries:
- The thesaurus entries have a topic ("Healthy" and "CN1CCCC1c2cccnc2," respectively). All entries on the page are related to this topic.
- Sub-topics are not necessarily "subsets" of (completely encompassed by) the main topic. For example, the drug "NICORETTEPLUS" might include other active or inactive compounds not encompassed by the main topic.
- Each sub-topic has information (data) associated with it that may or may not pertain directly to the main topic, yet all of the topics are related in a fundamental way.
- Data can be attached to the main topic of the thesaurus entry as well as the sub-topics.
- Several of the sub-topics have alternate meanings. For example, "Sound" might also appear in a thesaurus entry under "Noise," a meaning entirely separate from the one used above.

The following concepts are critical to understanding the basic TDT format:
The datafield is the fundamental block of information in a TDT. A datafield is simply a string of printable characters. In the case above, examples of datafields are: nicotine and C10H14N2. If a datafield contains one or more of the characters "$<>;|", it must be quoted. For example: "$10.50". Semicolons are used to separate individual datafields on a single line such as in 1.17;1 and 1.42;2.
A tag is a label that names the type of data in the datafield. A tag consists of one or more alphanumeric characters. i.e., A-Z, a-z, and 0-9, plus the underscore character "_". Tags that start with a dollar sign indicate an identifier such as $SMI and $CAS in the example above. Tags that start with a slash ('/') are non-identifiers that are automatically cross-referenced to the tree root which may or may not be a SMILES. All other tags indicate items that are data only. For example: PCN and MF
A dataitem consists of a tag followed by angle-bracket-enclosed datafields. No space is allowed between the tag and the opening "<" Examples of dataitems are: PCN<nicotine> and $CASPC<54-11-5>.
A datatype is a set of definitions that indicate the meaning of a dataitem's fields. Each datatype is identified by its tag. See the section below for a more complete description and examples.
A complete datatree is represented as a series of dataitems with at least one identifier and terminated with a '|' (pipe).
Additional examples of TDTs are:
-
$SMI<CCC>MF<C3H8>$CAS<123-45-6>BP<123.4>$ISM<[13CH2]C>BP<124.3>|
$CAS<765-43-2>XCD<92.8;99.2;98.3;collected at STP>|
$SMI<c1ccccc1>$WLN<R>PCN<benzene>|
There is no restriction to the length of a TDT, the length of an identifier, the length of any dataitem, or the length of any datafield. THOR provides mechanisms for storing "binary data", data that can include anything at all. However, in order to take advantage of the datatree structure of a TDT, it must be rooted in $SMI. Although, a TDT can be rooted in another identifier, it cannot contain mulitple branches.
7.5 Datatypes
A THOR datatype gives meaning to the data in a THOR database. For example, the data "$AID<1234-5> is meaningless without the definition of $AID, which tells us that it is an vendor catalog number. This is the primary purpose of datatype definitions.
A secondary function for datatype definitions is standardization of data and identifiers. Standardization is the process of modifying data according to particular instructions. For example, the datatype $NAM (name) is standardized by converting all characters to uppercase, eliminating all spaces and tabs, and removing punctuation. This greatly improves the likelihood that you will find what you are looking for. If, for example, you request "1,2-dichlorobenzene", THOR will retrieve a datatree that was entered even if it originally contained "1-2-DichloroBenzene"; both the stored name and your request will be converted to "12DICHLOROBENZENE". Note that some information is lost in this process, but carefully designed standardizations can be extremely helpful without sacrificing information.
7.5.1 Creating Datatypes
Datatype definitions are expressed as TDTs. This requires that there be at least a minimal set of predefined (truly universal) datatypes.
The following datatypes are the default datatypes:
| $D<tag> | Internal tag for datatype |
| _V<vtag[;vtag...]> | Name for label |
| _B<btag[;btag...] | Brief name for pull-down menu |
| _N<ntype[;ntype...]> | Normalization parameters |
| _P<[*][;[*]...]> | Merlin in-memory pool inclusion flag --- * for all or if number N, then at most N datafields will be loaded of that type; alternatively, a '!' creates a row in Merlin from each subtree rooted by the identifier datatype to which it applies |
| _S<summary> | One-line summary of datatype's meaning and use |
| _D<description> | Long description of datatype's meaning and use. |
| _M<set> | Membership of the datatype for pull-down submenus |
| _C | General comments |
| _O | Owner of the datatype |
Normally, a dataitem has a fixed number of fields. However, since these special datatypes are used to define the datafields of other datatypes, the _V specification can contain any number of datafields (each of which defines a datafield). Exactly one _V specification must occur in each datatype definition; it also sets the maximum number of datafields that may appear in the _N, _B, and _P specifications.
Here is a simple example that defines a SMILES datatype:
- $D<"$SMI">
-
_V<SMILES>
_B<SMILES>
_N<USMILES AUTOGEN $GRF>
_P<*>
_S<SMILES, primary identifier in the Daylight system>
_D<SMILES is the primary key to all data in THOR system>
_M<IDENTIFIER,SYSTEM,POOL>
_C<SMILES, the fundamental identifier in THOR databases>
_O<daylight@daylight.com (Daylight CIS, Inc., Irvine, CA>|
A more complex example is the definition of CLOGP, the computed partition coefficient of a compound. It has three fields in its definition (the real CLOGP is more complex):
- $D<CP>
-
_V<"CLOGP;#ERROR LEV;VERSION">
_B<"clogp;clogp/err;clogp/ver">
_P<"*;*;*">
_N<"REAL32;INDIRECT $I;">
_S<Estimate of the LogP(o/w) coefficient by CLOGP3>
_D<Estimate of the partition coefficient LogP(o/w)
computed by the CLOGP3 algorithm (from the MedChem
Project)>
_M<MODEL RESULTS,MEDCHEM,PHYSICAL PROPERTY,POOL>|
7.5.2 Standardization
The standardizations below work in a straightforward fashion: They modify the string representation of a single field in a straightforward fashion. During standardization of a TDT, if the THOR system discovers that the root of the TDT is not a SMILES, it will search the TDT for any datafield with a SMILES normalization. If one is found, it will use it to create a SMILES root for the TDT, "demoting" the original root to a subtree.
-
USMILES -- generate unique SMILES
ASMILES -- generate absolute SMILES (unique isomeric SMILES)
USMILESANY -- generate unique SMILES, unrelated to root
ASMILESANY -- generate absolute SMILES, unrelated to root
WHITE0, WHITE1, WHITE2 -- remove all, 2 or more, or 3 or more spaces, respectively
UPCASE -- convert to upper case
DOWNCASE -- convert to lower case
NOPUNCT -- remove punctuation
CASNUM -- insert hyphens and verify checksum
INDIRECT -- designate indirect data field
INTEGER16, INTEGER32, REAL32, REAL64 -- designate numeric data format
BINARY -- denote binary data
SMILES_NTUPLE n -- designate SMILES-order n-tuple data where n is the
n-tuple order; maintain order of data such as pairs of numbers
for 2D coordinates (e.g. SMILES_NTUPLE 2) so that there is a
one-to-one correspondence between the data and the molecule's atoms
even after other normalizations like generating unique SMILES
PART_NTUPLE n -- designates component-order n-tuple data where n is the number of data per part;
maintain order for datatypes like FPP with a set of N fingerprints
corresponding with N dot-disconnected SMILES representing a mixture or library
AUTOGEN tag -- generate new dataitem using contents of dataitem specified by 'tag'
GRAPH -- convert SMILES to GRAPH when retrieving data
MAKEGRAPH -- produce a GRAPH subtree of datatype $GRF
D3D -- compute 3D hash for use with $3D3 datatype
7.6 Database Anatomy
A THOR chemical database, usually thought of as a single entity, is actually made up of as many as three databases to store: datatypes, indirect references, and chemical information. By convention, these databases are referred to respectively as the datatype-definition database, the indirect-data database, and the regular database.
- datatypes
- The datatypes-definitions database contains the definitions of all datatypes. Datatypes are frequently common to all databases at a particular site (i.e. all regular databases refer to the same datatypes database); this makes the data from all databases intercompatible.
- indirect data
-
The optional indirect-data database contains the
expansions for indirect data. Like datatypes databases,
indirect-data database are often shared by several related
regular databases. Such indirect references save space when a
field's contents is repeated many times in a database and
allow uniformity by predefining a set of choices for users.
Indirect datafields are defined by adding the INDIRECT specification to the normalizations. When the TDT is retrieved from the database, the indirect reference is looked up in the indirect-data database and the expansion data replaces the original indirect reference.
-
chemical data
- The regular, or chemical-information database, contains data about chemicals.
7.7 Database Files
Each database actually consists of six files; these are described below:
- description
- The description file (suffix .THOR, also called the header file) describes the database. It contains the names of the other files, timestamps for each of the constituent files, the database's encrypted passwords, and the names of the databases where datatypes and indirect data can be found.
- lockfile
- A lockfile (suffix .LCK) is present whenever a THOR server has the database open. It contains the process ID of the THOR server, and prevents two THOR servers from opening the same database.
- primary data
- The primary data file (suffix .DP) contains all of the data in the database; this is where THOR datatrees are stored. A database can be completely rebuilt from the contents of this file.
- primary hash
- The primary hash file (suffix .HP) contains the hash table that allows "order one" (constant time) access of the primary data via SMILES.
- cross ref.
- The cross-reference file (suffix .DX) contains a cross-reference for each non-SMILES identifier; each non-SMILES identifier is listed with the SMILES of each TDT in which the non-SMILES identifier appears.
- cross-ref. hash
- The cross-reference hash file (.HX) contains the hash table that allows "order one" access of non-SMILES identifiers.
7.8 Reactions in THOR
Given that the THOR system uses SMILES strings for all of its database operations it works with reactions without modification. One can create a database with only molecules, a combination of molecules and reactions, or reactions only. It should be noted though that all reaction SMILES must be quoted within a datatree because of the ">" symbol used to separate the reactants and products within the SMILES string.
In many cases, combining molecule and reaction datatrees in a single database is the most appropriate organization for the database. When one builds a database, there is often a temptation to store molecule data (especially for the products) with the reaction. For example, physical data for reaction product(s) is commonly stored under the reaction. One perspective is that the physical data relates to the reaction because purification methods, solvents, etc. may affect the physical measurements and repeating the reaction multiple times will potentially give varying physical data. By the same token, the physical data can be considered to be about the product molecule and should be stored with it. The correct approach depends on the goals of the database and the types of queries which the end users wish to answer. Within the THOR system, both approaches are valid and may be employed.
There is one new and two extension of existing database standarizations that relate specifically to reaction processing.
-
MAKERXNMOL -- generate component molecules for a reaction;
parses dataitems into dot-separated components
PART_NTUPLE n -- designate SMILES-order ntuple data over the reaction components
SMILES_NTUPLE n -- designate SMILES-order ntuple data while ignoring ">" symbols
Go To Next Chapter... 8. Merlin - Exploratory Data Analysis Program