![]()
![]()
by
46 Uppergate Road, Stannington, Sheffield S6 6BX, UK
Tel. +44 144 233 3170; Fax +44 114 234 3415
The Daylight Chemical Information System includes an efficient implementation of the Jarvis-Patrick non-hierarchical clustering method. The Daylight Clustering package is widely used, and can handle extremely large files, but recent research has suggested that hierarchical methods (in particular, Ward’s method) perform better in property-prediction studies[1], and in separating known actives and inactives[2]. They can also avoid the problems associated with large numbers of singletons, coupled with a few large heterogenous clusters, which sometimes occur with the Jarvis-Patrick method.
This presentation describes a new version of BCI's Agglomerative Hierarchical Clustering Package, which has a number of features of potential interest to Daylight customers:
The diagram below shows a "clustering" of various different clustering methods[3-4]; those shown in green are those which have been implemented by BCI Ltd. In non-hierarchical methods, such as the Jarvis-Patrick method, all clusters are at the same level, whereas in hierarchical methods, larger clusters are subdivided into smaller ones, down to the level of individual compounds; a single "partition" (slice across the hierarchy) can then be taken at any level to give the desired number of clusters.
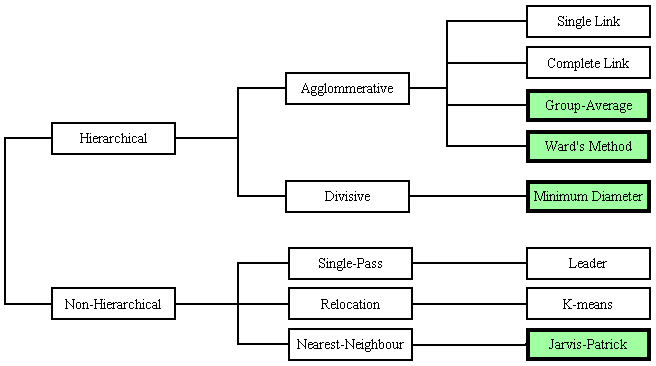
Agglomerative hierarchical clustering proceeds by a series of binary mergers (agglomerations), initially of individual compounds, later of clusters formed at previous stages, and the different methods are distinguished by the way in which the distance between clusters is measured, in order to choose the pair to be merged next. . In the two agglomerative methods implemented by BCI, Ward’s Method and the Group-Average Method, the distance measures are respectively the squared Euclidean Distance and the Cosine Coefficient, measured between the cluster "centroids", which are calculated as a weighted average of the descriptive features of the compounds in each cluster.
BCI's implementation uses the efficient Reciprocal Nearest Neighbour algorithm[5] whose time requirements vary with the square of the number of compounds being clustered (rather than with the cube, as in traditional implementations). Its space requirements are linear with the number of compounds, rather than varying with the square. This allows quite large files to be processed in reasonable time: on a Silicon Graphics Challenge, a file of 100,000 compounds can be clustered in about 54 cpu hours.
BCI's package supports input and output of TDT files, and thus provides a useful alternative, or adjunct, to Daylight's own Jarvis-Patrick implementation. The programs are able to read fingerprint (FP<>) data directly from TDT files, and to add CL<> data items for a user-selected number of clusters to output files, which can then be processed by Daylight’s ListClusters and ShowClusters programs, or using the ClusterView viewing program available in the Daylight "contributed programs" directory. No toolkit routines are involved in BCI's programs, and thus Daylight licenses are not required to run them.
An option for the output of a hierarchical cluster membership file allows the Hierarchical Cluster Viewer program (also available in the Daylight "contributed programs" directory) to be used; this is discussed below.
BCI’s package is able to cluster compounds on the basis of either binary structure fingerprints, or continuous-varying property data (for example, topological indexes or calculated physicochenical properties). In the former case, the fingerprints may be supplied either as a Daylight TDT file containing fingerprint data, or as a BCI-format file derived from BCI’s MAKEBITS fingerprint-generation program. In the latter case, either "raw" or normalised property values for each compound can be supplied in a simple text file format; the programs are able to normalise raw data (to avoid biases introduced by the greater numerical range of certain property values) over either the range of each variable, or over their standard deviations. Missing values for individual compounds are handled by ignoring the variable in question for that compound.
In addition to adding CL<> data items to Daylight TDT files, a separate output file can be generated in which the structure identifiers for the members of each cluster are sorted by position in the original file, alphanumerically, or by increasing distance from the cluster centroid. A file showing the compounds in clusters at all levels of the hierarchy can also be generated.
Two programs are included in the Agglomerative Hierarchical Clustering Package:
The following diagram shows the processing flow involved:
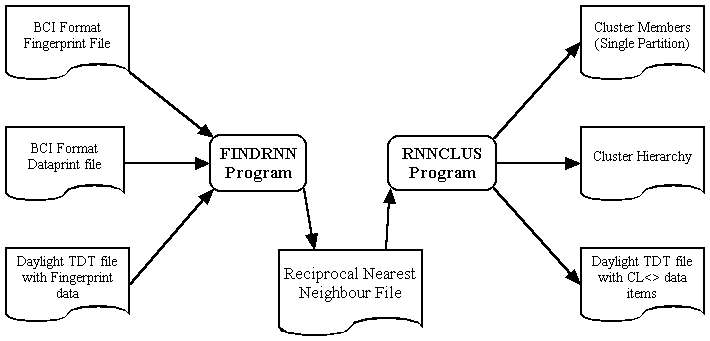
Both programs have a simple command line interface, and are available for either UNIX or PC implementation. There is no fixed limit on the maximum number of objects which can be clustered.
This program was written by Dr Robert Brown of Abbott Laboratories, specifically for purpose of viewing the hierarchical clusters obtainable from BCI's clustering programs. It is freely available from the Daylight Contributed Programs directory, though it is a Daylight Toolkit program, and requires a Daylight toolkit license to run. Thanks are due to Rob for contributing the program, and for providing the illustrations shown below.
HierView requires as input the BCI-format hierarchical cluster membership file, and a file listing the SMILES for the clustered compounds in the same order as they appeared in the fingerprint file used for clustering. The program is then able to display two clusters at a time (several pages are used when the cluster has too many compounds to be shown on one screen). These are clusters which merge together further up the hierarchy, and by using control buttons to move up and down the hierarchy, comparing the contents of the clusters merged at each stage, the user can decide the best level at which to "draw a line" across the hierarchy and extract a single partition. Slider bars in the control window can also be used to jump around the hierarchy.
Here, relatively low down the hierarchy, we have two clusters, one with two members, and one with three.

These clusters will merge together higher up the hierarchy, and the merged cluster can be shown by clicking on the Merge A and B button, which leads to the following screen, in which it appears as Cluster B. Returning to the previous screen can be acheived by clicking on the Divide B button.
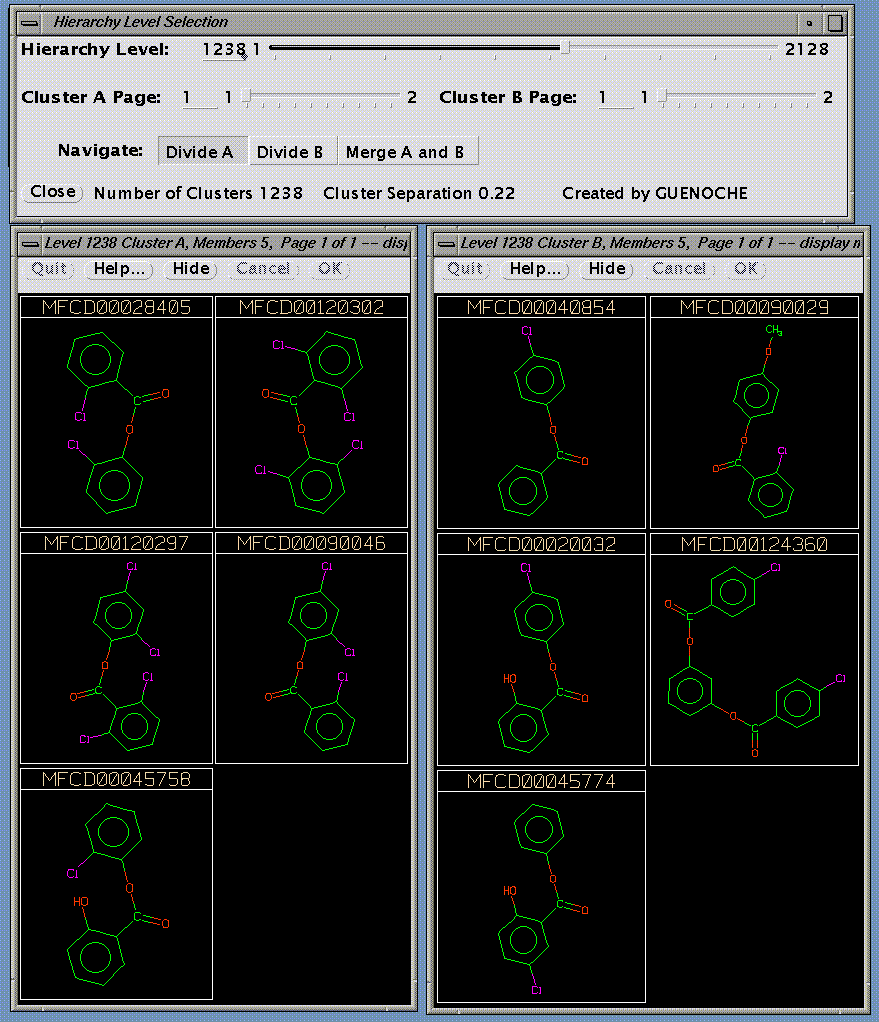
Similarly, the above two clusters can be merged, giving cluster A in the next screen.

Such mergers can be continued until the top of the hierarchy is reached, with just two clusters containing all the compounds in the dataset. Successive division operations will eventually lead to singleton clusters at the bottom of the hierarchy.
For further information on BCI's Agglomerative Hierarchical Clustering Package, and on other BCI products and services, please contact:
| Dr John M. Barnard Barnard Chemical Information Ltd. 46 Uppergate Road Stannington Sheffield S6 6BX UK |
Dr Geoff M. Downs Barnard Chemical Information Ltd. 30 Kiveton Lane Todwick Sheffield S31 0HL UK |
| Tel.: +44 114 233 3170 Fax: +44 114 234 3415 Email: barnard@bci1.demon.co.uk |
Tel.: +44 1909 515444 Fax: +44 1909 774074 Email: downs@bci2.demon.co.uk |
or visit our Website at http://www.bci1.demon.co.uk.